It is well known that the transmission of malaria is dependent on temperature, and of particular importance is how temperature controls the longevity of mosquitoes of the Anopheles genus. The temperature driven survival of these mosquitoes, is one of the main causes of the current global distribution of malaria.
The relationship between temperature and malaria transmission has made it possible to construct climate based malaria maps of the theoretical distribution of malaria, dividing severely affected areas, from areas with occasional epidemics. The same methodologies have also been used to project the future impact of climate change on malaria.
In 2013 two scientific papers were published describing how many of these models have been wrong headed. One by Mordecai and colleges, and one by our research group at Centre for International Health and Bjerknes Centre for Climate Research.Both papers claim malaria is transmitted most efficiently around 25 degrees Celsius, 6 °C lower than previous models. Roughly this means places with temperatures below or over 25 °C theoretically will have a lower potential for efficient spread of malaria compared to areas where temperatures are around 25 °C.
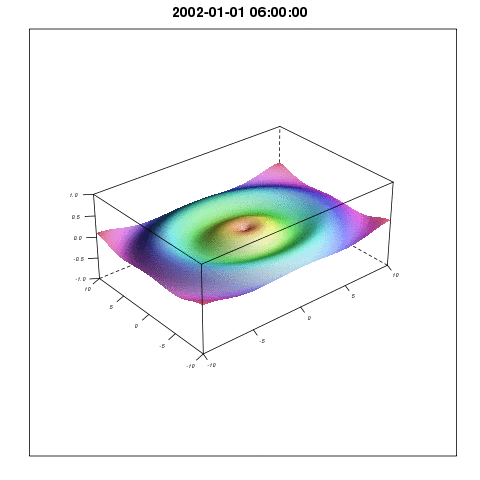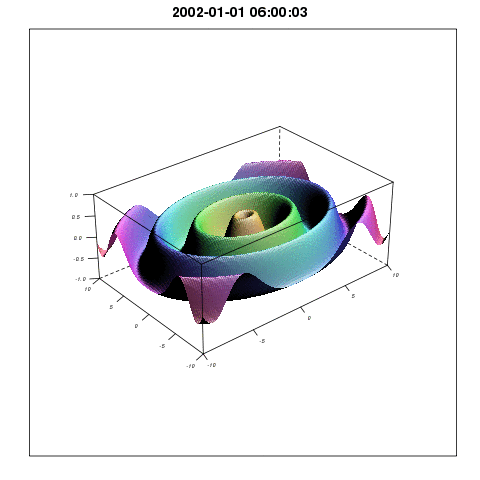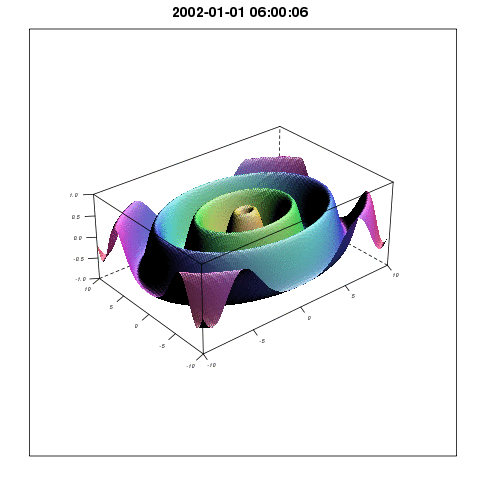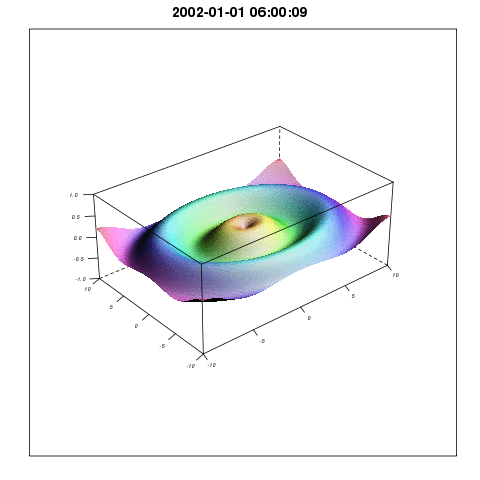
To describe the relationship between temperature and mosquito mortality, researchers use theoretical models. One model has been particular popular among scientists, and has been used in for example MARA/ARMA model used by the World Health Organization, to model the global constraints of malaria , and to project the consequences of global warming on malaria here, and here. In our paper we show that this model is not supported by observations, and there are reasons to believe that either the results shown in the mentioned papers are correct for the wrong reasons, or that the results themselves are wrong.
Researchers should now rethink how climate has influenced malaria in the past, present and future.